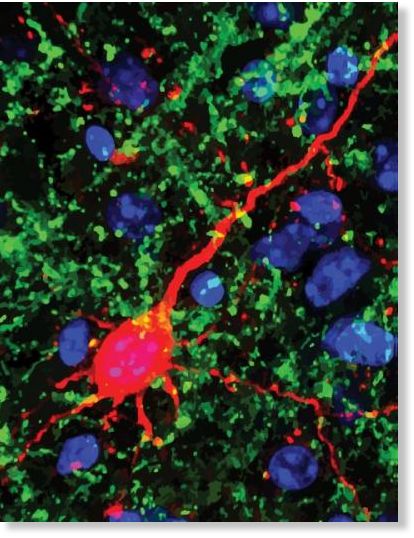
© Emily Petrus and Amal IsaiahWhen adult mice were kept in the dark for about a week, neural networks in the auditory cortex, where sound is processed, strengthened their connections from the thalamus, the midbrain's switchboard for sensory information. As a result, the mice developed sharper hearing. This enhanced image shows fibers (green) that link the thalamus to neurons (red) in the auditory cortex.
Call it the Ray Charles Effect: a young child who is blind develops a keen ability to hear things that others cannot. Researchers have long known that very young brains are malleable enough to re-wire some circuits that process sensory information. Now researchers at the University of Maryland and Johns Hopkins University have overturned conventional wisdom, showing the brains of adult mice can also be re-wired to compensate for a temporary vision loss by improving their hearing.
The findings, published Feb. 5 in the peer-reviewed journal
Neuron, may lead to treatments for people with hearing loss or tinnitus, said Patrick Kanold, an associate professor of biology at UMD who partnered with Hey-Kyoung Lee, an associate professor of neuroscience at JHU, to lead the study.
"There is some level of interconnectedness of the senses in the brain that we are revealing here," Kanold said.
"We can perhaps use this to benefit our efforts to recover a lost sense," said Lee. "By temporarily preventing vision, we may be able to engage the adult brain to change the circuit to better process sound."
Kanold explained that there is an early "critical period" for hearing, similar to the better-known critical period for vision. The auditory system in the brain of a very young child quickly learns its way around its sound environment, becoming most sensitive to the sounds it encounters most often. But once that critical period is past, the auditory system doesn't respond to changes in the individual's soundscape.





Comment: