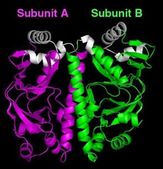
The NIST/BNL team reports that it has defined - for the first time - the structure of a "metabolic switch" found inside most types of bacteria - the cyclic AMP (cAMP) receptor protein, or CRP - in its "off" state. CRP is the "binding site" (attachment point) for cAMP, a small molecule that, once attached, serves as the signal to throw the switch. This "on" state of CRP then turns on the genes that help a microbe survive in a human host.
The researchers hope that once the switching mechanism is understood the data can be used to develop new methods for preventing tuberculosis and other pathogenic bacterial diseases.




